
#NVIDIA CUDA TOOLKIT DOCUMENTATION DOWNLOAD#
You can also download the installer to your local computer and upload it to the GPU instance.
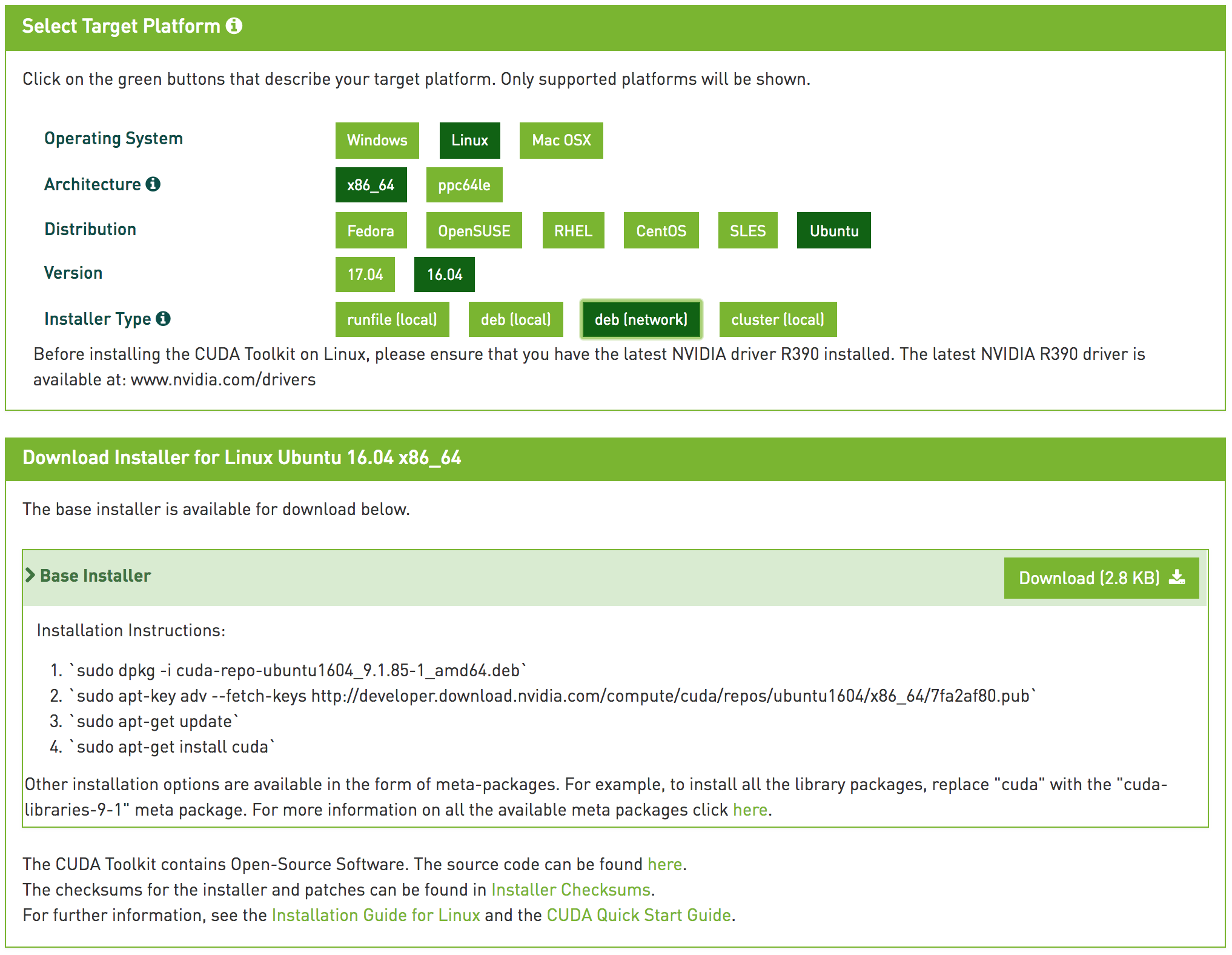
Go to the CUDA Toolkit download page or visit.
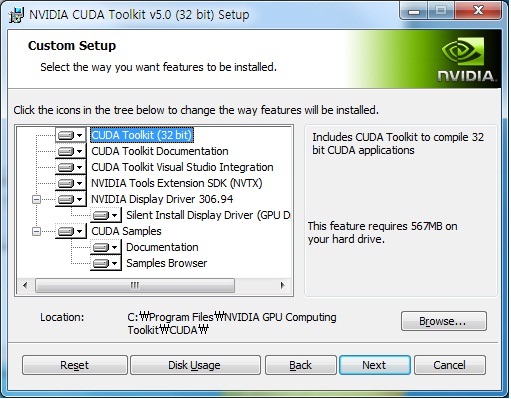
Installing CUDA Toolkit on a Linux instance
#NVIDIA CUDA TOOLKIT DOCUMENTATION HOW TO#
This document uses the most common CUDA Toolkit 10.1 as an example to describe how to install CUDA Toolkit on a GPU instance. The compiled programs can be run on CUDA-enabled processors.īecause GPU instances use NVIDIA graphic cards, you must install the CUDA Toolkit. The CUDA platform is designed to work with programming languages such as C, C++, and Fortran. It includes the CUDA instruction set architecture (ISA) and the parallel computing engine within the GPU. With a generic parallel computing architecture, CUDA allows GPUs to solve complex computing problems. The new ASE-3.0 Sphinx page is now up and running! ()Ī beta version of the new ASE-3.Compute Unified Device Architecture (CUDA™) is a computing platform developed by NVIDIA. Ten people fromĬAMd/Cinf will do a “doc-sprint” from 9 to 16. Thursday April 24 will be ASE documentation-day. Possibility to calculate Infrared intensities (13ĪSE version 3.0.0 released (13 November 2008).Īsap version 3.0.2 released (15 October 2008).Īn experimental abinit interface released (9 June 2008). Improved ase.vibrations module: More accurate and Web-page now uses the Read the Docs Sphinx Theme (20 February 2016). The Atomic Simulation Environment | A Python library for working withĪSE version 3.13.0 released (7 February 2017).ĪSE version 3.12.0 released (24 October 2016).ĪSE version 3.10.0 released (17 March 2016). get_potential_energy () -31.492847800329216 Supported Calculators ¶ĪCE-Molecule amber DeePMD-kit DMol³ Gaussian Grimme DFT-D3 gulp Mopac qmmm tip3p ~deMon-NanoĪSE version 3.22.1 released (1 December 2021).ĪSE version 3.22.0 released (24 June 2021).ĪSE version 3.21.1 released (24 January 2021).ĪSE version 3.21.0 released (18 January 2021). calc = NWChem ( xc = 'PBE' ) > opt = BFGS ( h2 ) > opt. # Example: structure optimization of hydrogen molecule > from ase import Atoms > from ase.optimize import BFGS > from import NWChem > from ase.io import write > h2 = Atoms ( 'H2'.
